Novel 3D imaging model may show path to more water-efficient plantsA new computational pipeline for analyzing three-dimensional imaging data can...
Published on by Water Network Research, Official research team of The Water Network
A new computational pipeline for analyzing three-dimensional imaging data can help biologists more accurately and quickly see how the cells in a plant's leaves respond to the environment and identify plants that more efficiently use water, according to researchers.
A team of computer scientists and biologists from Penn State developed a 3D imaging model to study how tiny structures called stomatal guard cells, which are involved in plant photosynthesis and transpiration, interact with neighboring cells when undergoing physical changes.
The model is more efficient and accurate than existing methods of analyzing cellular geometry and mechanics, and the researchers found that the guard cells behaved in unexpected ways. The research will help biologists run experiments more efficiently and identify plants, including important agricultural crops, that can better adapt to a changing climate.
"Currently, it takes experts five to eight hours to manually label just the guard cells in a single 3D image set," said Dolzodmaa Davaasuren, a doctoral candidate in Penn State's College of Information Sciences and Technology who led development of the pipeline. "Our team wanted to automate processes so we could study more images."
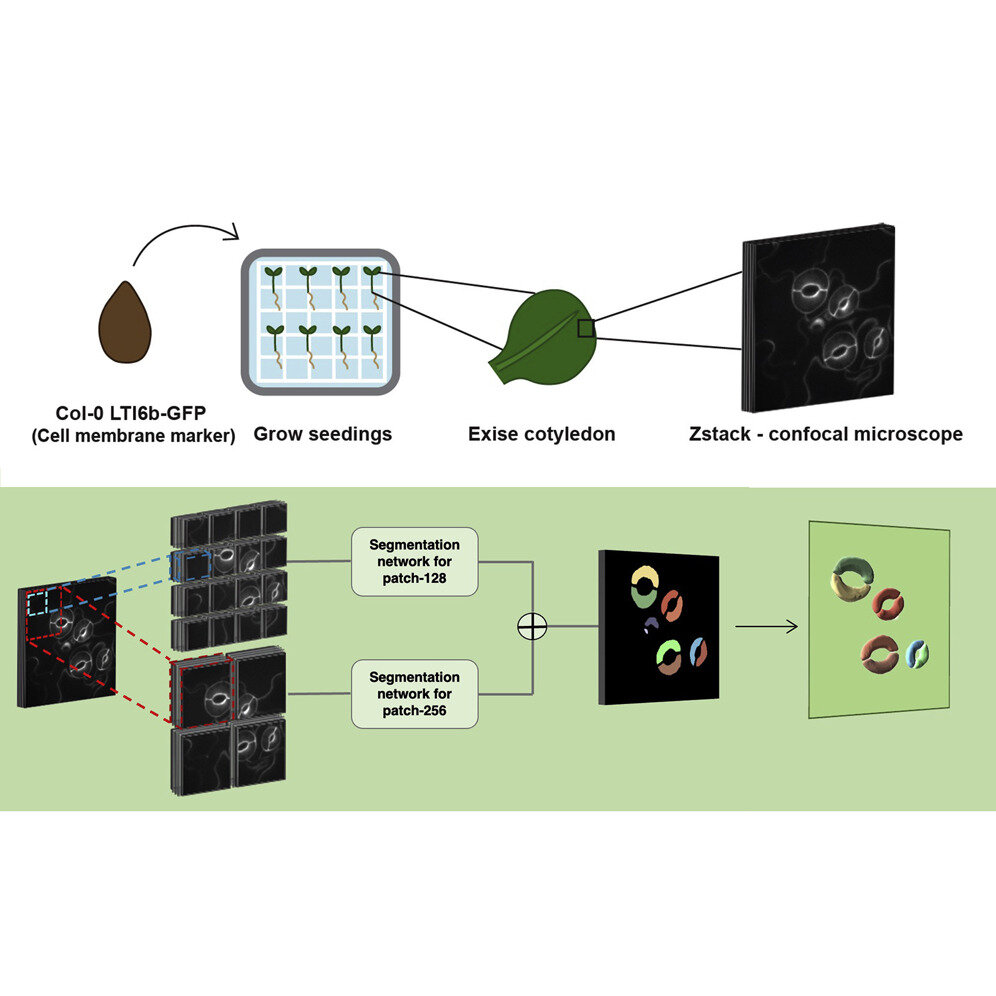
The researchers built and tested their pipeline using the model plant Arabidopsis thaliana, commonly known as thale cress. They used a specialized confocal microscope to take 3D images of guard cells on the leaves of the plant. Guard cells surround stomatal pores and regulate how much carbon dioxide and water vapor pass through the pores. The team collected images before and after ablating, or using a laser beam to poke holes in, neighboring cells that were touching guard cells to see how stomatal volume changed.
The scientists used the 3D U-Net segmentation model as a basis for their model, which they called 3D CellNet, and added an encoder that better preserves spatial information. They also added an attention module, which tells the model to focus on specific parts of the 3D image. In this case, they told the module to focus on the tiny guard cells. The researchers used just five manually labeled 3D images to train their model. Further image processing steps were taken in the pipeline to measure the shapes of the guard cells.
The team found that their new pipeline labeled images and measured cell volumes more quickly and accurately than trained cell biologists. They also found that 3D CellNet segmentation outperformed the base model on which it was built plus two additional 2D models. They reported their findings in the journal Patterns.
Attached link
https://phys.org/news/2022-12-3d-imaging-path-water-efficient.htmlTaxonomy
- Sustainable Agriculture
- Sustainable Agriculture
- Agriculture
- Regenerative Agriculture